Bioinformatics
The Centre for Computational Biology champions a multidisciplinary approach to research and teaching in Data Sciences for Life Sciences. Our researchers link the fields of Mathematics, Statistics, and Computer Sciences to Biology and Medicine, while reaching a broad community of data scientists on campus. We lead the development of novel integrative and analytical approaches to data starting with Omics and Clinical.
Our Data Sciences experts have partnered with scientists and healthcare professionals of Chengdu’s West China Hospital – one of China’s biggest and best hospitals – to help link innovative ‘Omics’ data with clinical information for a range of conditions. This collaborative working is vital to help transform the lives of patients, as well as understanding the cause of diseases.
Theme Co-Leads

Professor Jean-Baptiste Cazier
Chair of Bioinformatics
Director of the Centre for Computational Biology
View profile
 Professor Georgios Gkoutos
Professor Georgios Gkoutos
Chair of Clinical Bioinformatics
View profile
Research groups
Spotlight on research using drug structure analysis to identify new therapeutics for breast cancer
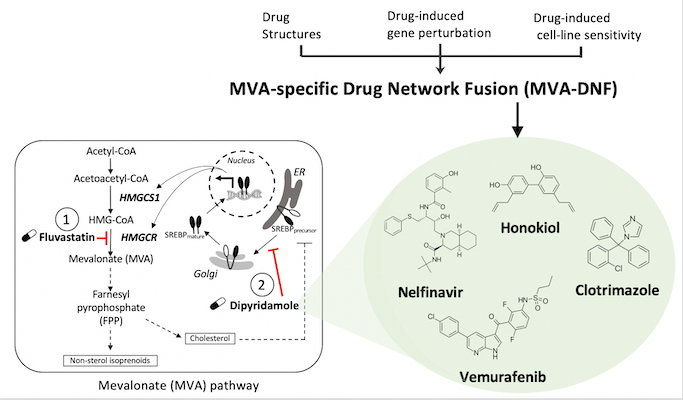
In a study led by Deena Gendoo’s lab, an integrative pharmacogenomics pipeline was developed to identify novel and effective therapeutics for breast cancer therapy. This pipeline, mevalonate drug-network fusion (MVA-DNF), integrated drug-induced gene perturbation signatures, drug sensitivity, and drug structure profiles to generate a pathway-centric network of drugs that can target the mevalonate (MVA) pathway.
The mevalonate pathway is a metabolic pathway that is a hallmark of many cancer types, including breast cancer, as it produces cholesterol and other isoprenoids that are essential for cell proliferation and survival. Dipyridamole (DP) is a drug which synergizes with statins to potentiate the pro-apopotic activity of statins as part of the mevalonate pathway, but this drug has limited clinical utility for cancer patients. Using MVA-DNF, new statin-drug combinations were identified that target the MVA pathway, and these hits have been validated by testing on cancer cell-lines and patient-derived organoids, showing their potential for breast cancer therapy.
Accordingly, this work identified several actionable compounds and novel and effective therapeutics to help combat difficult-to-treat triple-negative breast cancer.
van Leeuwen J.E., Ba-Alawi W., Branchard E, Cruickshank J, Schormann W, Longo J, Silvester J, Gross P, Andrews D, Cescon D, Haibe-Kains B, Linda P, Gendoo D. Computational pharmacogenomic screen identifies drugs that potentiate the anti-breast cancer activity of statins. Nat Commun 13, 6323 (2022).
Spotlight on COVID-19 and Leukaemia
New study describes which cancer patients are more vulnerable to COVID-19.
Lancet Oncology 21: 1309-1316 (2020). Lee LYW, JB Cazier, T Starkey, SEW Briggs, R Arnold, V Bisht, . . . UKCCMP Team. COVID-19 prevalence and mortality in patients with cancer and the effect of primary tumour subtype and patient demographics: a prospective cohort study.
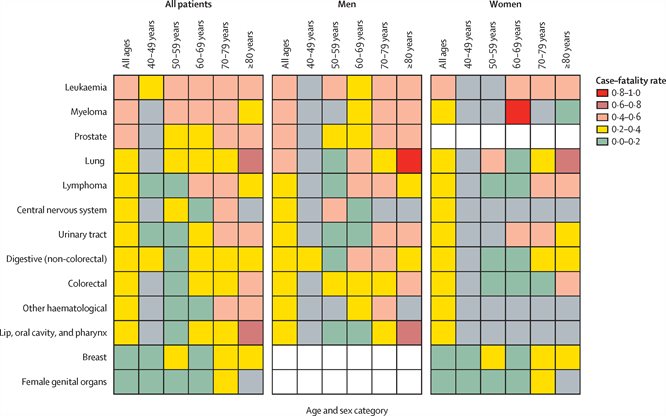
A newly published study involving Lennard Lee and Jean-Baptiste Cazier has found that, compared to other cancers, patients with blood cancers are more vulnerable to the effects of the coronavirus pandemic. The researchers were able to determine that patients with haematological cancers, particularly older patients and those with leukaemia, had a more severe COVID-19 trajectory compared to patients with solid organ tumours.
Selected Highlights from this Research Theme
Eur Heart J (2023). Gill SK, A Karwath, HW Uh, VR Cardoso, Z Gu, A Barsky, . . . GV Gkoutos, D Kotecha, BigData@Heart Consortium and the cardAIc group. Artificial intelligence to enhance clinical value across the spectrum of cardiovascular healthcare. doi: 10.1093/eurheartj/ehac758
JAMA Netw Open 5:e220130 (2022). Varnai C, C Palles, R Arnold, HM Curley, K Purshouse, VWT Cheng, . . . J-B Cazier, The UKCCMP Team. Mortality Among Adults With Cancer Undergoing Chemotherapy or Immunotherapy and Infected With COVID-19.
Cancer Med Online: doi: 10.1002/cam4.4941 (2022). Bahcivanci B, R Shafiha, GV Gkoutos and A Acharjee. Associating transcriptomics data with inflammatory markers to understand tumour microenvironment in hepatocellular carcinoma.
Lancet 398:1427-1435 (2021). Karwath A, KV Bunting, SK Gill, O Tica, S Pendleton, F Aziz, . . .GV Gkoutos, D Kotecha, The cadrdAlc group and the Beta-blockers in Heart Failure Collaborative Group. Redefining beta-blocker response in heart failure patients with sinus rhythm and atrial fibrillation: a machine learning cluster analysis.
Cell 184:4612-4625 e4614 (2021). Almarri MA, M Haber, RA Lootah, P Hallast, S Al Turki, HC Martin, . . . C Tyler-Smith. The genomic history of the Middle East.
mSystems 6:e0034621 (2021). Benkwitz-Bedford S, M Palm, TY Demirtas, V Mustonen, A Farewell, J Warringer, . . . D Moradigaravand. Machine Learning Prediction of Resistance to Subinhibitory Antimicrobial Concentrations from Escherichia coli Genomes.
Eur J Obstet Gynecol Reprod Biol 261:211-216 (2021). Craciunas L, O Pickering, J Chu, M Choudhary, J Zurauskiene and A Coomarasamy. The transcriptomic profile of endometrial receptivity in recurrent miscarriage.
Cancers 13:2779 (2021). Dragomir I, A Akbar, JW Cassidy, N Patel, HW Clifford and G Contino. Identifying Cancer Drivers Using DRIVE: A Feature-Based Machine Learning Model for a Pan-Cancer Assessment of Somatic Missense Mutations.
mSphere 6 e00738-20 (2021). Murphy R, M Palm, V Mustonen, J Warringer, A Farewell, L Parts and D Moradigaravand. Genomic Epidemiology and Evolution of Escherichia coli in Wild Animals in Mexico.
American J Human Genetics 107:149-157 (2020). Haber M, J Nassar, MA Almarri, T Saupe, L Saag, SJ Griffith, . . . C Tyler-Smith. A Genetic History of the Near East from an aDNA Time Course Sampling Eight Points in the Past 4,000 Years.
Nat Commun 11:1825 (2020). Chung PED, DMA Gendoo, R Ghanbari-Azarnier, JC Liu, Z Jiang, J Tsui, . . . E Zacksenhaus. Modeling germline mutations in pineoblastoma uncovers lysosome disruption-based therapy.
Am J Hum Genet 107:149-157 (2020). Haber M, J Nassar, MA Almarri, T Saupe, L Saag, SJ Griffith, . . . C Tyler-Smith. A Genetic History of the Near East from an aDNA Time Course Sampling Eight Points in the Past 4,000 Years.
Lancet Oncol 21:1309-1316 (2020). Lee LYW, JB Cazier, T Starkey, SEW Briggs, R Arnold, V Bisht, . . . UKCCMP Team. COVID-19 prevalence and mortality in patients with cancer and the effect of primary tumour subtype and patient demographics: a prospective cohort study.
Cancer Cell 38:306-307 (2020). Lee LYW, T Hill, O Topping, M Tilby, M Baker, J Greig, . . . UKBCCC- Project. Utility of COVID-19 Screening in Cancer Patients.
Front Genet 11:669 (2020). Mountford HS, P Villanueva, MA Fernandez, L Jara, Z De Barbieri, LG Carvajal-Carmona, . . . JB Cazier, DF Newbury. The Genetic Population Structure of Robinson Crusoe Island, Chile.
Sci Rep 9:17405 (2019). Althubaiti S, A Karwath, A Dallol, A Noor, SS Alkhayyat, R Alwassia, . . . GV Gkoutos, R Hoehndorf. Ontology-based prediction of cancer driver genes.
PLoS Comput Biol 15:e1006596 (2019). Gendoo DMA, RE Denroche, A Zhang, N Radulovich, GH Jang, M Lemire, . . . B Haibe-Kains. Whole genomes define concordance of matched primary, xenograft, and organoid models of pancreas cancer.
PLoS Comput Biol 14:e1006258 (2018). Moradigaravand D, M Palm, A Farewell, V Mustonen, J Warringer and L Parts. Prediction of antibiotic resistance in Escherichia coli from large-scale pan-genome data.
Genome Med 9:119 (2017). Moradigaravand D, T Gouliouris, B Blane, P Naydenova, C Ludden, C Crawley, . . . SJ Peacock. Within-host evolution of Enterococcus faecium during longitudinal carriage and transition to bloodstream infection in immunocompromised patients.
BMC Bioinformatics 17:140 (2016). Zurauskiene J and C Yau. pcaReduce: hierarchical clustering of single cell transcriptional profiles.
Nat Commun 5:3756 (2014). Cazier JB, SR Rao, CM McLean, AK Walker, BJ Wright, EE Jaeger, . . . FC Hamdy. Whole-genome sequencing of bladder cancers reveals somatic CDKN1A mutations and clinicopathological associations with mutation burden.