Research Theme within School of Biosciences:Molecular and Cell Biology
Birmingham Fly Facility: https://www.birmingham.ac.uk/research/birmingham-fly-facility/birmingham-fly-facility.aspx
RNA processing and translation
Gene expression, the fundamental process that decodes the genetic blueprint, is particularly complex in eukaryotes. The primary pre-mRNA transcripts undergo several covalent modifications in the nucleus before the resulting mature mRNAs are exported to the cytoplasm. Nuclear processing includes 5' end capping, splicing and 3' end processing/polyadenylation. In the cell these processes are interconnected and coupled to transcription (1). In addition, by affecting nuclear mRNA-protein (mRNP) complexes formation, RNA processing can also have a regulatory role on protein synthesis, influencing mRNA export, mRNA localization, translation and mRNA degradation (1).
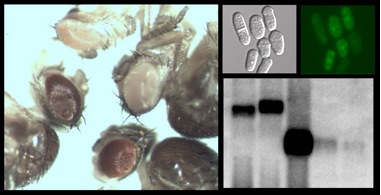
We are interested in several aspects of RNA processing, and we study them primarily in the yeast Schizosaccharomyces pombe and the fruitfly Drosophila melanogaster because of their genetic advantages. One of our main concerns is the mechanism of nonsense mediated mRNA decay (NMD) - a translation coupled process that plays an important role in quality control of gene expression by degrading abnormal mRNAs carrying premature termination codons (PTCs) (2,3). Our interest in NMD has been ignited by observations that PTCs affect spliced mRNAs much more than transcripts from intron-less genes (2); and by our finding that PTCs can affect nuclear polyadenylation (2,4). These observations imply the existence of a link between pre-mRNA processing, translation and mRNA stability. By studying NMD we hope to better understand the mechanisms of these interconnections.
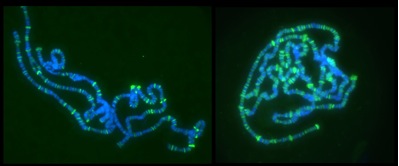
A second concern of our research is the development of techniques to visualize the recruitment of processing factors to nascent transcripts. Nascent RNAs are decorated with numerous factors, forming complex RNPs; the composition/conformation of these RNPs commits the pre-mRNAs to particular maturation pathways and determines the fate of the mature mRNAs (e.g., affect NMD). To visualize nascent RNPs we use the polytene chromosomes of D. melanogaster the pictures above, shows active Pol II (left) and Sm proteins (right), DNA in blue. In these expanded interphase chromosomes, each locus contains about a thousand copies of each gene and so they provide an effective system in which to analyze the association of individual proteins with sites of transcription and the nascent pre-mRNA. We track specific factors either by immunostainig or by using transgenic flies expressing fluorescently tagged proteins.
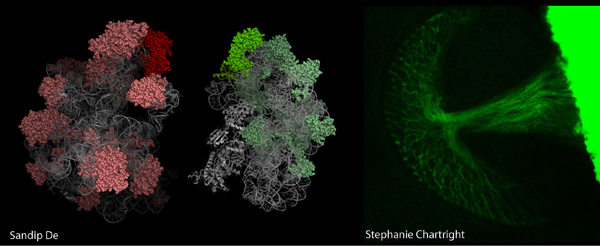
A third interest of the group is to visualize translation. We have tagged the small and large ribosomal subunits with fluorescent proteins (see picture above). We use this technique to visualize ribosomal subunits interaction sites in Drosophila cells, including neurons (the picture above shows the fly optical nerve expressing a GFP-tagged ribosomal protein). By using this technique we have made the remarkable discovery that translating ribosomes are also present in the cell nucleus (5). We are now interested in identifying the ribosome or ribosomal subunits-associated RNAs within the nucleus.
Applications for PhD student and postodoc positions are welcome at any time.
References
- Moore, M. J., and Proudfoot, N. J. (2009) Cell 136(4), 688-700
- Brogna, S., and Wen, J. (2009) Nat Struct Mol Biol 16(2), 107-113
- Muhlemann, O., Eberle, A. B., Stalder, L., and Zamudio Orozco, R. (2008) Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms 1779(9), 538-549
- Brogna, S. (1999) RNA 5(4), 562-573
- Al-Jubran et al.(2013). RNA 19(12):1669-83.
The lab has received funds from The Royal Society and Wellcome Trust.