
Professor Aga Gambus
Professor of DNA Metabolism
Staff profile for Professor Aga Gambus, Professor of DNA Metabolism, Department of Cancer and Genomic Sciences, College of Medicine and Health, University of Birmingham


Problems during DNA duplication (DNA replication) are thought to be a major cause of mutations that are observed in cancer. Our research focuses, therefore, on understanding the organization of protein machinery involved in DNA replication and its regulation especially through modification by small modifiers: ubiquitin and SUMO.
Our cells contain about 2 metres of DNA that needs to be faultlessly duplicated before every cell division to produce two identical daughter cells. Amazingly, most of the time our cells manage to achieve this task. Sometimes mistakes do happen and, if not repaired efficiently, they can lead to mutations and genomic instabilities. DNA replication therefore plays a major role in the development of cancer, genetic diseases and aging.
DNA replication is one of the most fundamental processes in life and thus is very highly conserved throughout evolution. This fact facilitates meaningful studies of the DNA replication process in model systems. In our laboratory we use Xenopus laevis egg extract system which is the only higher eukaryotic cell-free system capable of efficient replication of template DNA in vitro and thus provides an invaluable model system for biochemical studies of DNA replication. Once we discover novel ways of regulation of replication process using Xenopus system, we translate our finding into human cells using immortalised human cell lines.
We use these systems to study the organization and regulation of the DNA replication machinery. In particular we are interested in understanding how modification of replication factors by attachment of small protein modifiers (ubiquitin and SUMO) regulates this process. Ubiquitylation and sumoylation are essential tools for modulating the function of proteins involved in the response to DNA damage. A series of specialised enzymes: E1, E2 and E3 covalently attach single or multiple ubiquitin or SUMO moieties to target proteins most often via lysine residues. All of the seven lysine residues of ubiquitin can be used for further ubiquitylation resulting in the formation of ubiquitin chains. In addition, there are three forms of SUMO (1,2 and 3). All these create a wide scope for the functional complexity of ubiquitin and SUMO signalling. The research in our group focuses on understanding how these modifications regulate the process of eukaryotic DNA replication and damage.
We have discovered recently the first elements of the mechanism of unloading of the replication machinery (replisome) once it fulfilled its essential role. Our findings provided the first insights into the process of termination of eukaryotic DNA replication.
To ensure faultless duplication of the whole genome, DNA replication initiates from thousands of origins of replication. An origin fires when the replicative helicases are activated and start to unwind double stranded DNA creating two DNA replication forks. The progressing replication forks move through the chromatin until they encounter forks from the neighbouring origins. When the forks converge the termination of replication forks happen: the helicases pass each other, synthesis of DNA is completed, the replisomes disassemble and topisomerase II resolves the daughter DNA molecules (see model below).
We have identified the first elements of the mechanism of replisome disassembly at the termination of eukaryotic DNA replication and the role of ubiquitylation in this process. We found that only one subunit of the replicative helicase, Mcm7, is polyubiquitylated at termination and that following this ubiquitylation, the helicase is removed from chromatin in a p97/VCP segregase-dependent manner. As the helicase forms the organising centre of the replisome, its removal leads to disassembly of the remaining replication machinery (Moreno et al. Science 2014). Subsequently, we identified the Cul2Lrr1 ubiquitin ligase as the enzyme ubiquitylating Mcm7 in higher eukaryotes (Sonneville et al. NCB 2017). Finally, any replisomes that are not unloaded in S-phase are unloaded in mitosis in a back-up pathway driven by activity of TRAIP ubiquitin ligase (Priego Moreno et al. Life Sci Alliance 2019).
We believe that disruption of the replisome disassembly process is detrimental to human health and that understanding the basis of this process will provide valuable targets that could be therapeutically exploited. We have, however, barely begun to understand the mechanism behind this process. Our efforts now focus, therefore, on determining the detailed mechanism and regulation of this process both in Xenopus laevis egg extract and in human cells. We aim also to understand what are the consequences of disruption of this process.
Activation and regulation of TRAIP ubiquitin ligase (funded by BBSRC)
We have shown that TRAIP ubiquitin ligase interacts with post-termination replisome in S-phase but it is efficiently ubiquitylating Mcm7 within the replisome only in mitosis (Priego Moreno et al. Life Sci Alliance 2019). Interestingly, Mcm7 in ubiquitylated with K63 or K6 chains in mitosis and these types of chains can lead to mitotic replisome disassembly.
Our aim is to understand how TRAIP is activated and regulated to productively ubiquitylate Mcm7 only in mitosis and only when replication fork approaches DNA damage during S-phase.
The role of SUMO in regulation of chromosomal replication
The process of sumoylation resembles closely this of ubiquitylation, but the role of SUMO modification during DNA replication and damage are much less understood. We aim therefore to identify replication factors modified my SUMO during the process of replication and characterise the function of these modifications.
SinMolTermination will develop a real-time and super resolution single-molecule imaging platform (Figure) to monitor the progression and termination of replication forks and directly observe protein dynamics within the replisome.
Using this platform we aim to:
Discover how misregulation of termination can give rise to replication stress.
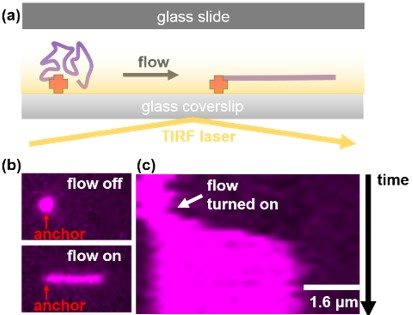
Figure: Combining microfluidics and TIRF microscopy to visualise and manipulate single DNA molecules. (a) DNA molecules that are anchored inside a microfluidic flow cell can be extended using hydrodynamic flow and imaged using TIRFM. (b) Micrograph of a fluorescently stained DNA molecule where flow is turned on and off. Notice the molecule elongates under flow. (c) Kymograph of the DNA molecule shown in (b) showing that the molecule rapidly elongates when flow is turned on.
Gambus A. Termination of Eukaryotic Replication Forks. Adv Exp Med Biol. 2017;1042:163-187. doi: 10.1007/978-981-10-6955-0_8. PMID:29357058
Sonneville R., Priego MorenoS., Knebel A., Johnson C., Hastie C.J., Gartner A., Gambus A. and Labib K. CUL-2LRR-1 and UBXN-3 drive replisome disassembly during DNA replication termination and mitosis. Nat Cell Biol. 2017 Apr 3. doi: 10.1038/ncb3500. Joined corresponding author.
We greatly enjoy sharing our enthusiasm for science with members of the public. We believe that it is possible to get across very complex ideas of cell biology – sometimes this involves simplifying the terminology, building models and drawing on analogies: anything that allows people to access these concepts and to share our excitement for discoveries.
Apart from opening our lab to members of the public we also take part in fundraising activities and university community days.
 |
 |
Above left: DNA repair race game. Birmingham University Community Day 2012. Above right: Cancer therapy game. Birmingham University Community Day 2013.
 |
 |
Above Left: Frog cake for the Great Science Cake Off competition 2013: to explain our science through the art of baking! Above right: Helping the fundraising. CRUK Relay for life 2013.
 |
|
Above Left: Scale model of a nucleus created for Cancer Showcase It has 200km of thread inside it. Above Right: Model of DNA replication fork made of wine gums – educational and yummy. Made for Cancer Showcase 2013.